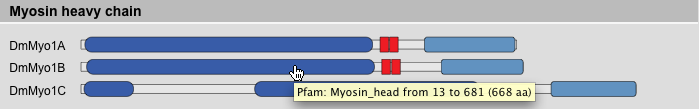
This result view provides the domain composition schemes of the selected sequences. The sequences are sorted in the following order: protein class, protein variant, taxonomy. This ordering should be best to compare the domain composition of homologous proteins, e.g. HsMyo1A, MmMyo1A, and GgMyo1A. Of course, this holds true only if the protein sequences are classified and named consistently. Tooltips provide information about domain names and coordinates.
