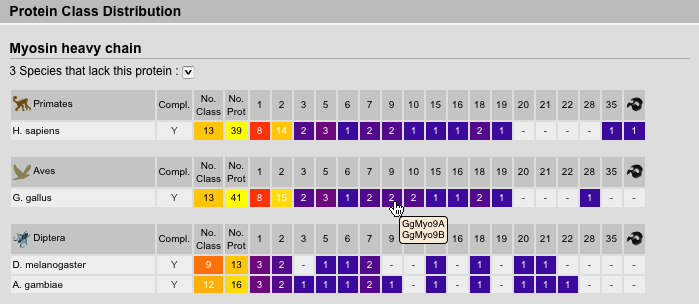
The protein inventory is sorted by proteins followed by taxonomy. Of course, this view is most useful, if all homologs of a protein were selected and not subclasses. Species names are abbreviated but full names are available via tooltips. The survey completeness as well as number of classes and sequences are given. A survey of the genome for a specific protein is thought to be complete if the corresponding genome sequence is almost finished (assembly with high covarage data available) and we have finished our in-depth analysis not being able to identify further homologs of the protein. Next the number of sequences corresponding to a certain class are given. Tooltips on these numbers show the list of sequence names.
